Welcome to MetaBrain
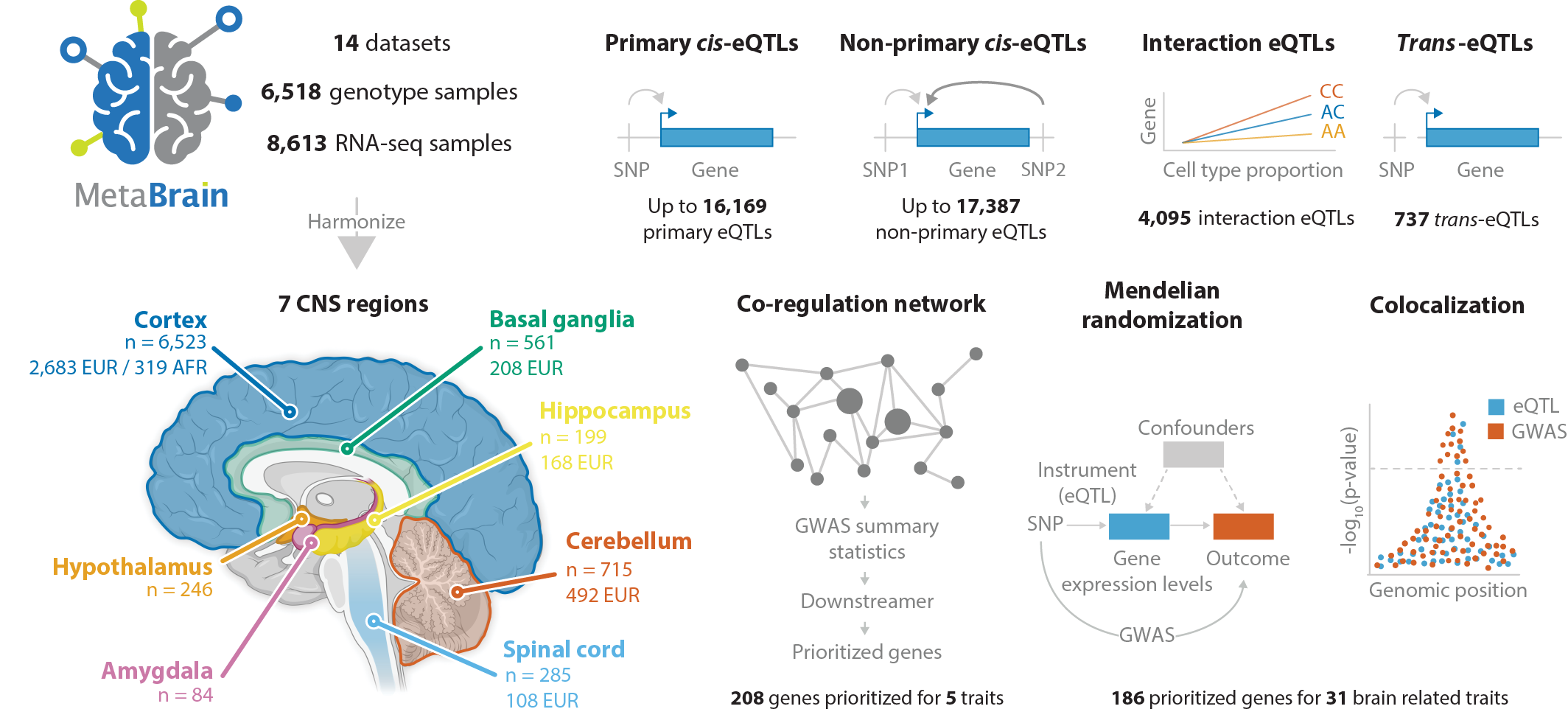
MetaBrain is a large scale eQTL meta-analysis of previously published human brain eQTL datasets. On this website you can find a browser for the cis-eQTL and trans-eQTL meta-analysis in Cortex samples from European ancestry. Additionally, this website provides a gene network that can be used to annotate genes and to perform gene set enrichment analyses.Overview of MetaBrain
Disclosure / Terms of use
By using this website or downloading these data, you and your collaborators (“investigators”) signify your assent to all of the following conditions:- These data are provided on an “AS-IS” and “AS-AVAILABLE” basis for scientific research and educational use only; without warranty of any type, expressed or implied (including but not limited to any warranty as to their performance, merchantability, or fitness for any particular purpose); and without warranty as to the accuracy, completeness, currency or reliability of any content available through this website.
- Use of this website and the content available on this website is at investigators’ sole risk. Investigators are responsible for taking all necessary precautions to ensure that any content you may obtain from the website is free of viruses.
- MetaBrain results cannot be re-distributed for any purpose.
- Investigators certify that they are in compliance with all applicable local, state, and federal laws and/or regulations and institutional policies regarding use of this data (e.g., including regarding human subjects and genetics research).
- Investigators will cite the appropriate MetaBrain publication reference(s) when presenting or publishing on results based (directly or indirectly) on this data. (Please refer to references listed below.)
- Investigators will never attempt to identify any participant.
How to cite
Please Cite: Brain expression quantitative trait locus and network analysis reveals downstream effects and putative drivers for brain-related diseases by N. de Klein et al., Nature Genetics, 2023, doi: https://doi.org/10.1038/s41588-023-01300-6Contact
For questions, bug reports, or requests, please contact Ellen Tsai (ellen.tsai@biogen.com); Lude Franke (ludefranke@gmail.com); Niek de Klein (niekdeklein@gmail.com); or Harm-Jan Westra (h.j.westra@umcg.nl).Summary statistics
Top effects can be browsed by using the links in the bar above (cis-eQTLs, trans-eQTLs). Full cis-eQTL summary statistics can be downloaded after filling in this form, which asks your name, e-mail address, institute name, and a short description of the intended use. A link to the downloads will then be sent to the email addres you provided. If you have any issues with the form, if you don't receive the download link, or if you have issues with the downloads themselves, please contact us.Update 2023-01-13: We have updated the cis-eQTL summary statistics on the download website following updated analysis procedures.
Update 2023-07-13: Summary statistics for the trans-eQTL and ieQTL analysis are available as supplementary tables 17 and 8 (respectively) with our paper. You can download them from Nature Genetics. Please contact us if you have specific requests regarding summary statistics.